Posted by star
on 2019-08-14 23:13:12
Hits:534
Scientists have identified two enzymes that protect chromosomes from oxidative damage and shortening. Blocking cancer cells could be a new strategy for treating cancer. Their role may be to prevent telomerase from doing its job, leaving chromosomes unprotected, and dividing chromosomes eventually disappearing. But telomeres contract over time, so a cell's lifespan is not infinite. Telomere shortening is the cause of cell aging.
Scientists have identified two antioxidant enzymes that work together to prevent the oxidation of DNA at the ends of chromosomes. In addition, in an experiment, scientists destroyed two enzymes associated with the structure of telomeres in cancer cells.
Now, scientists think a promising target for new cancer treatments is telomerase. In general, telomerase prevents telomere shortening in germ cells and stem cells, which helps with development. But telomerase also keeps their telomeres intact in cancer cells, keeping them in a constant state of division. Effective blocking of telomerase is therefore key, but the attempt is still in its infancy. The discovery of cooperative enzymes opens up new opportunities for indirect blocking of telomerase. "Instead of inhibiting the enzyme itself, it can turn to its substrate, making the ends of chromosomes unable to extend telomerase,” the scientists said.
Posted by star
on 2019-08-13 19:25:26
Hits:553
When RNA viruses attack the cells, the RIG-I-like receptors or Toll-like receptors sense pathogen-associated molecular patterns, and signal downstream through interferon regulatory factors (IRFs), transcription factors induce synthesis of type I and type III interferons. RNA viruses have evolved sophisticated strategies to disrupt these signalling pathways and evade elimination by cells. So it is important to know how IRFs maintain basal levels of protection against viruses.
The researchers from University of North Carolina at Chapel Hill and Tokyo Metropolitan Institute of Medicine depleted antiviral factors linked to RIG-I-like receptor and Toll-like receptor signalling to map critical host pathways restricting positive-strand RNA virus replication in immortalized hepatocytes and identified an unexpected role for IRF1. The results showed that constitutively expressed IRF1 acts independently of mitochondrial antiviral signalling (MAVS) protein, IRF3 and signal transducer and activator of transcription 1 (STAT1)-dependent signalling to provide intrinsic antiviral protection in actinomycin D-treated cells.
IRF1 localizes to the nucleus, where it maintains the basal transcription of a suite of antiviral genes that protect against multiple pathogenic RNA viruses, including hepatitis A and C viruses, Zika virus and dengue virus. The IRF1 protein in hepatocytes regulates RARRES3, an enzyme that attacks the virus when it is attempted to express in cells.
“Our findings reveal an unappreciated layer of hepatocyte-intrinsic immunity to these positive-strand RNA viruses and identify previously unrecognized antiviral effector genes.” the researcher said.
EIAAB SCIENCE INC, WUHAN has developed IRF1 protein, antibody and ELISA kit.
Welcome scientific research workers to choose and purchase.
EIAAB SCIENCE INC, WUHAN has developed
Posted by star
on 2019-08-13 00:36:51
Hits:525
Inflammation is an important component of the body's response to infection and tissue damage, but uncontrolled inflammation can lead to various diseases and promote cancer formation.
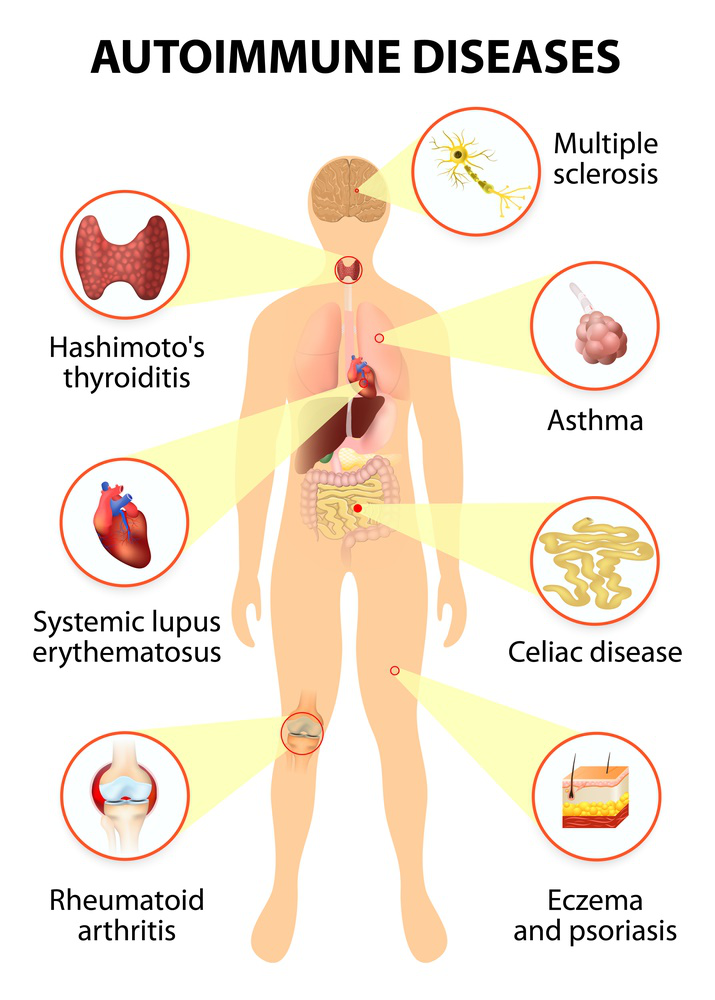
A new study led by the University of Texas MD Anderson Cancer Center found that a protein regulatory factor called Trabid is an important part of the autoimmune inflammation of the central nervous system in patients with multiple sclerosis.
The proinflammatory cytokines interleukin 12 (IL-12) and IL-23 connect innate responses and adaptive immune responses. They are also involved in autoimmune and inflammatory diseases. When the researchers deleted Zranb1 (which encodes Trabid) in dendritic cells, the induction of the expression of IL12 and IL23 by Toll-like receptors (TLRs) was inhibited. This impaired the differentiation of inflammatory T cells and protected mice from autoimmune inflammation. Trabid facilitated TLR-induced histone modifications at the promoters of IL12 and IL23, which involved deubiqutination and stabilization of the histone demethylase Jmjd2d.
“ Our findings highlight an epigenetic mechanism for the regulation of IL12 and IL23 and establish Trabid as an innate immunological regulator of inflammatory T cell responses.” the researchers said.
Trabid and Jmjd2d may be potential therapeutic targets for the treatment of inflammatory diseases such as multiple sclerosis.
EIAAB SCIENCE INC, WUHAN has developed Trabid and Jmjd2d protein, antibody and ELISA kit.
Welcome scientific research workers to choose and purchase.
Posted by star
on 2019-08-12 00:57:23
Hits:483
Under normal circumstances, the immune system recognizes and fights cancer cells, then eliminates them. However, sometimes this process can fail to form a tumor. Researchers at the Texas A&M Health Sciences Center found that when cancer cells block the function of the NLRC5 gene, they can escape the immune system and proliferate.
NLRC5 regulates major histocompatibility complex (MHC) class I genes. These genes encode molecules on the cell surface that present fragments of foreign proteins, such as proteins from viruses or bacteria. These protein fragments inform a portion of the immune system, called cytotoxic T cells, to trigger an immediate response from the immune system against this particular foreign antigen.
The new findings of this study showed the same system should strive to destroy cancer cells, but sometimes they can find a way to disable the NLRC5 gene, allowing them to escape the immune system and form tumors. "If MHC class I antigens do not work, cancer cells will not be killed by T cells," the researchers said. "We found that various mechanisms in cancer reduce the function and expression of NLRC5, and as a result, tumor cell escapes from the immunity system."
Biopsy results from 7747 solid tumor patients in The Cancer Genome Atlas (TCGA) database showed that the expression of NLRC5 gene is highly correlated with the survival rate of various cancer patients, especially melanoma, rectal cancer, bladder cancer, Cervical and head and neck cancers, patients with longer survival tend to have more NLRC5 expression.
"NLRC5 is an important biomarker for cancer. We can predict how long cancer patients can survive and how cancer treatment works for them." The researchers said: "NLRC5 can be a novel biomarker for cancer patients' survival and therapeutic response. It is also a potential target for new therapies."
EIAAB SCIENCE INC, WUHAN has developed NLRC5 protein, antibo......
Posted by star
on 2019-08-06 19:15:20
Hits:557
A study published in Cell reveals that vitamin D supplementation may slow the progression of type 2 diabetes because vitamin D improves glucose metabolism and helps prevent the development and progression of diabetes.
Type 2 diabetes is a chronic disease that places a huge burden on patients and society and can lead to serious health problems, including nerve damage, blindness and kidney failure. The risk of developing type 2 diabetes is often determined by several risk factors, such as obesity and family history. In some studies, there was no data linking low vitamin D levels to diabetes risk. In particular, these Numbers are small and not representative.
In the study, Matthew Smith, PhD, of Duke University, and colleagues looked at whether vitamin D supplementation had an effect on glucose metabolism in people at high risk for type 2 diabetes or diabetes. Insulin function and glucose metabolism were measured before and after vitamin D supplementation. Thirty-six percent of the participants had low levels of vitamin D at the start of the study, but when vitamin D supplementation was remeasured three months later, insulin function increased significantly.
At the same time, no improvement in glucose metabolism was detected, possibly because of improvements in metabolic function or longer treatment periods to see benefits.
Dr. Matthew Smith suggests that future studies should evaluate whether vitamin D supplementation is a response to individual factors. But before doing so, people at high risk for type 2 diabetes or diabetes should be advised to follow certain vitamin D supplements.